neuroConstruct 1.6.0
Free Version
Publisher Description
neuroConstruct is being developed in the Silver Lab in the Department of Neuroscience, Physiology and Pharmacology at UCL. neuroConstruct has been designed to simplify development of complex networks of biologically realistic neurons, i.e. models incorporating dendritic morphologies and realistic cell membrane conductances. It is implemented in Java and generates script files for the NEURON and GENESIS simulators, with support for other simulation platforms (including PSICS, MOOSE and PyNN) in advanced stages of development. It uses the latest NeuroML specifications, including MorphML, ChannelML and NetworkML.
Development of this software was made possible with funding from the Wellcome Trust, the Medical Research Council and the EU Synapse Project.
Some of the key features of neuroConstruct are:
* neuroConstruct can import morphology files in GENESIS, NEURON, Neurolucida, SWC and MorphML format for inclusion in single cell or network models, or more abstract cells can also be built manually.
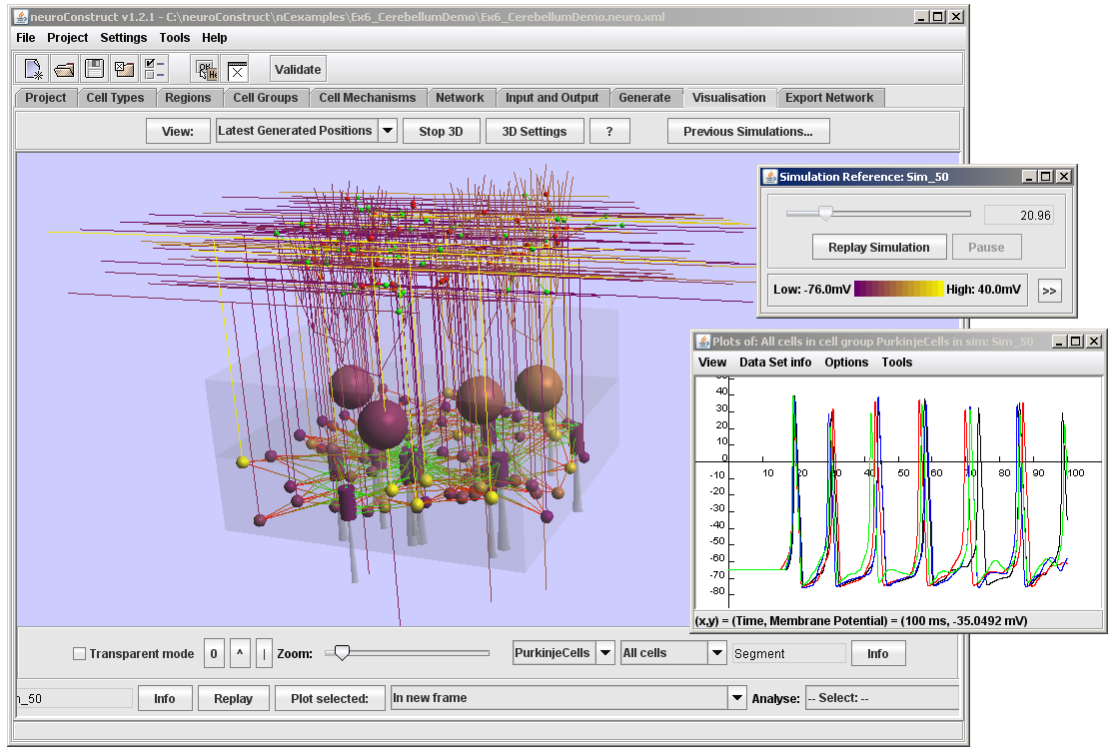
* Creation of networks of conductance based neurons positioned in 3D
* Complex connectivity patterns between cell groups can be specified for the networks
* Simulation scripts can be generated for NEURON, GENESIS, MOOSE, PSICS and PyNN based simulators (note: not every project can be generated for every simulator)
* Biophysically realistic cellular mechanisms (synapses/channel mechanisms) can be imported from native script files (*.mod or *.g) or created from templates using ChannelML
* Automatic generation of code to record simulation data and visualisation/analysis of data in neuroConstruct
* Recorded simulation runs can be viewed and managed through the Simulation Browser interface
* A Python based scripting interface can be used to control model generation and execution, allowing multiple simulations to be run for cell and network model optimisation
About neuroConstruct
neuroConstruct is a free software published in the Science list of programs, part of Education.
This Science program is available in English. It was last updated on 27 March, 2024. neuroConstruct is compatible with the following operating systems: Linux, Mac, Other, Unix, Windows.
The company that develops neuroConstruct is University College London. The latest version released by its developer is 1.6.0. This version was rated by 2 users of our site and has an average rating of 3.5.
The download we have available for neuroConstruct has a file size of 52.43 MB. Just click the green Download button above to start the downloading process. The program is listed on our website since 2012-08-23 and was downloaded 244 times. We have already checked if the download link is safe, however for your own protection we recommend that you scan the downloaded software with your antivirus. Your antivirus may detect the neuroConstruct as malware if the download link is broken.
How to install neuroConstruct on your Windows device:
- Click on the Download button on our website. This will start the download from the website of the developer.
- Once the neuroConstruct is downloaded click on it to start the setup process (assuming you are on a desktop computer).
- When the installation is finished you should be able to see and run the program.